Community Detection with the Map Equation and Infomap: Theory and Applications
Jelena Smiljanić, Christopher Blöcker, Anton Holmgren, Daniel Edler, Magnus Neuman, Martin Rosvall
ACM Computing Surveys 58(7), 1-34 (2026) [pdf] [arXiv:2311.04036]
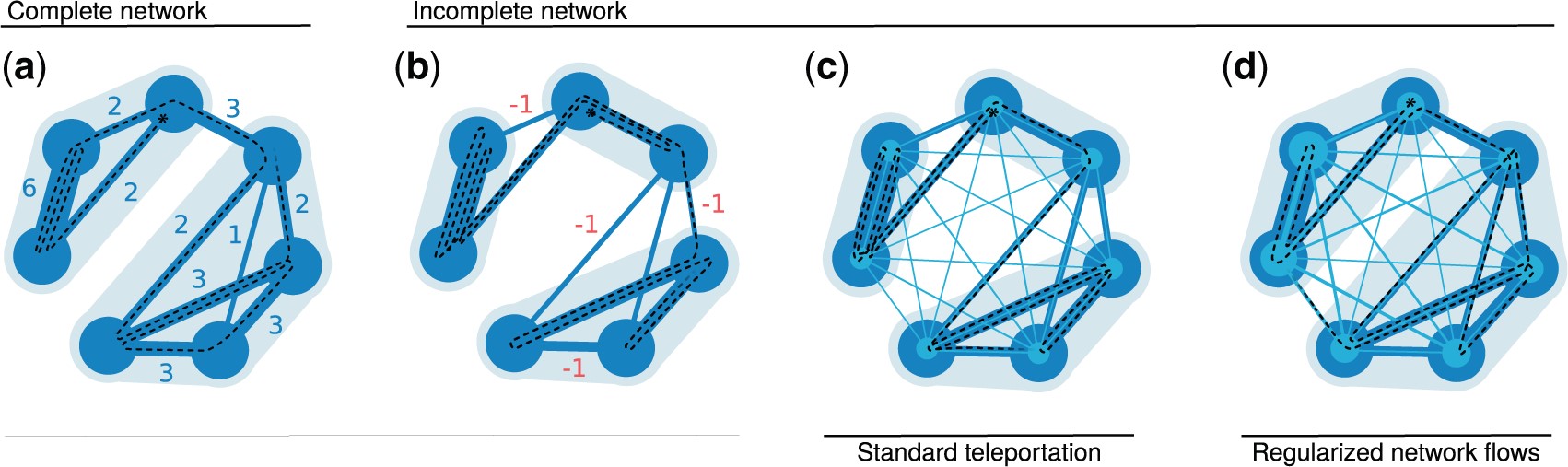
Real-world networks have a complex topology comprising many elements often structured into communities. Revealing these communities helps researchers uncover the organizational and functional structure of the system that the network represents. However, detecting community structures in complex networks requires selecting a community detection method among a multitude of alternatives with different network representations, community interpretations, and underlying mechanisms. This tutorial focuses on a popular community detection method called the map equation and its search algorithm Infomap. The map equation framework for community detection describes communities by analyzing dynamic processes on the network. Thanks to its flexibility, the map equation provides extensions that can incorporate various assumptions about network structure and dynamics. To help decide if the map equation is a suitable community detection method for a given complex system and problem at hand — and which variant to choose — we review the map equation’s theoretical framework and guide users in applying the map equation to various research problems.
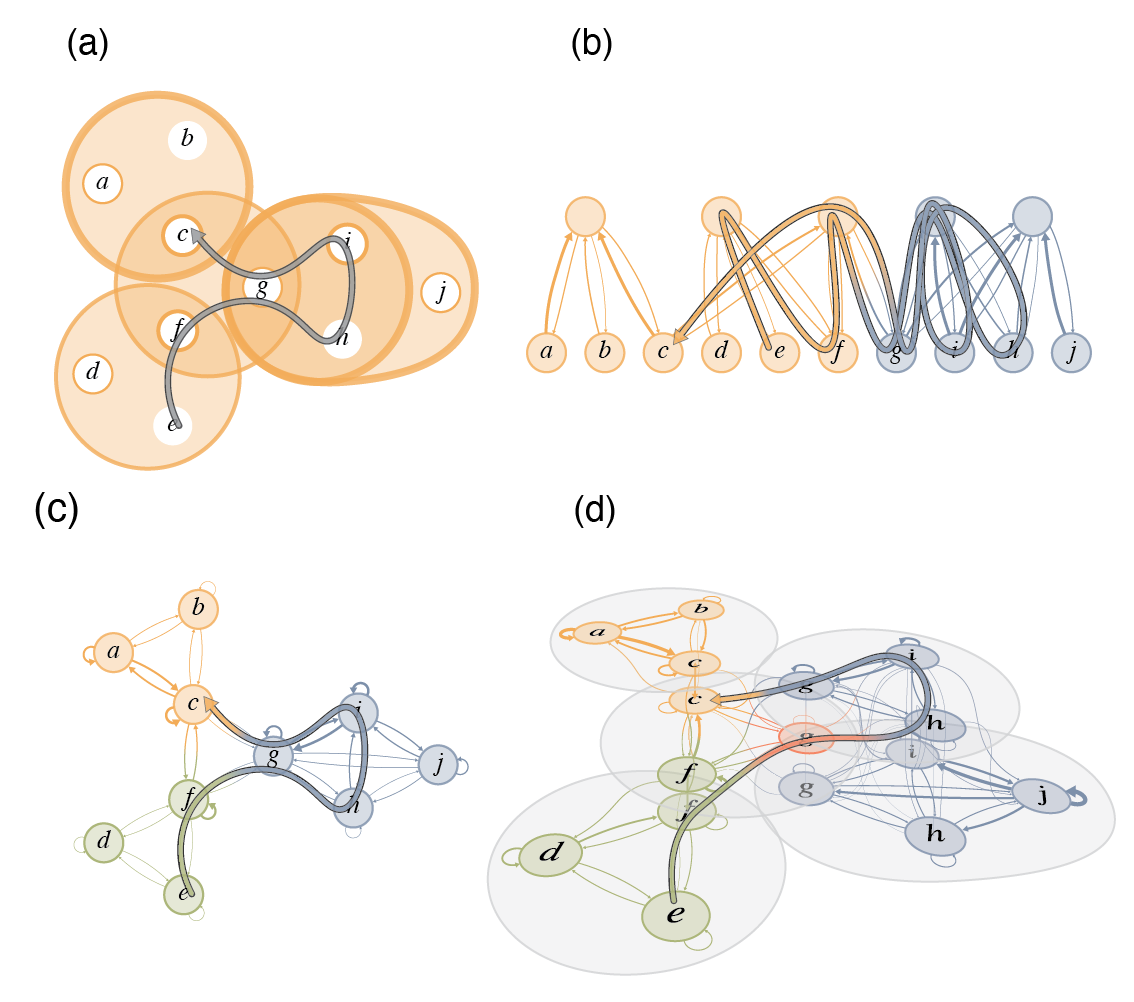
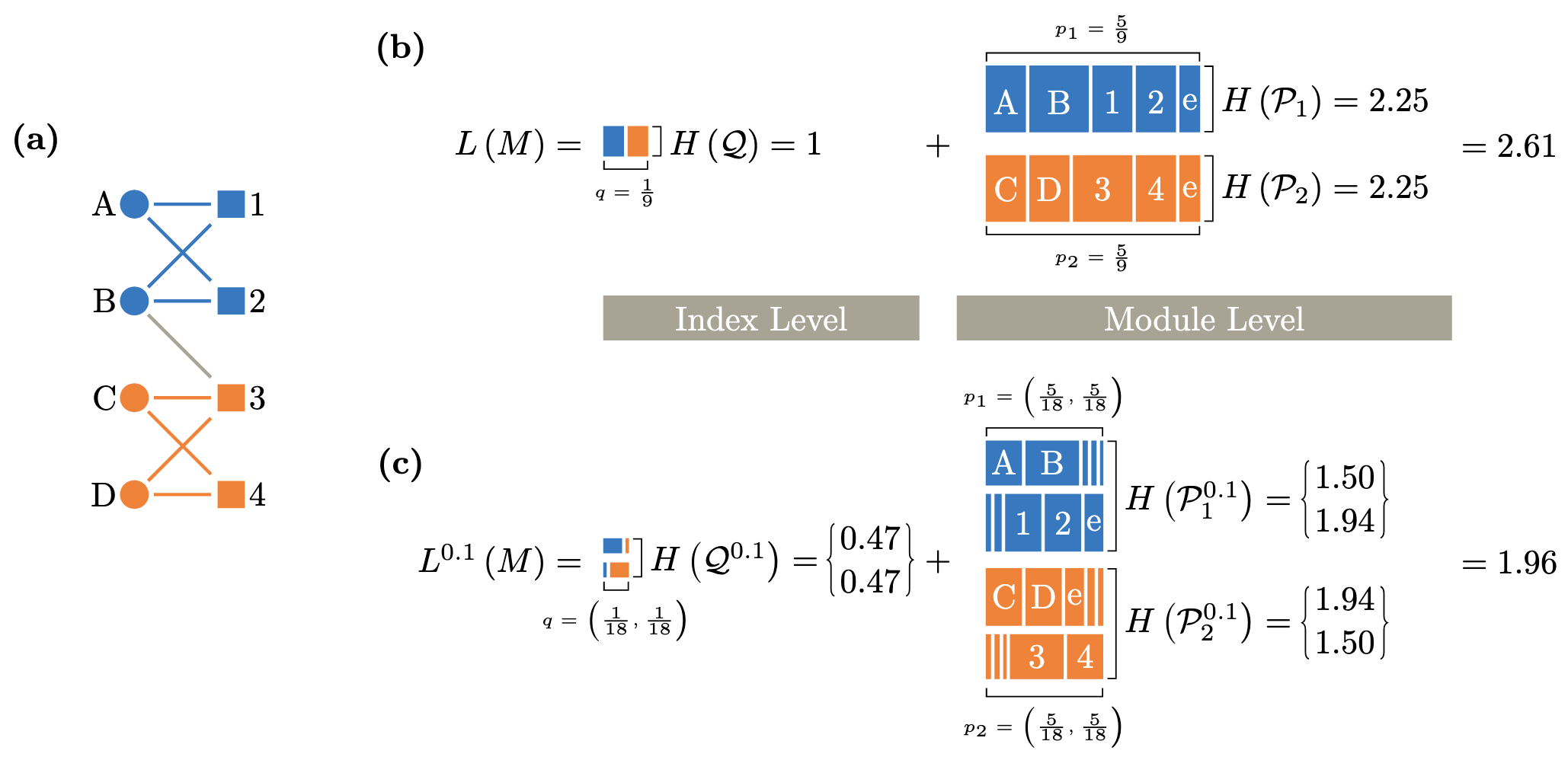
![FIG. Lazy encoding random walks on a schematic network with metadata. In this case, the random walk always encodes the next step if the target node's metadata is the same as the current node. When the metadata differs, the walker encodes with probability p in [1/3, 1] in a, p in [1/10, 1/3) in b, and p < 1/10 in c. Node shapes represent metadata, and node colors represent optimal partitions. Random walks with labeled circles where they encode and dashed lines where they do not, colored by the currently encoded module.](/assets/img/lazy-random-walk.png)